Applying multimodal biological foundation models across therapeutics and patient care

Healthcare and life sciences decision making increasingly relies on multimodal data to diagnose diseases, prescribe medicine and predict treatment outcomes, develop and optimize innovative therapies accurately. Traditional approaches analyze fragmented data, such as ‘omics for drug discovery, medical images for diagnostics, clinical trial reports for validation, and electronic health records (EHR) for patient treatment. As a result, decision makers (CxOs, VPs, Directors) often miss critical insights hidden in the relationships between data types. Recent advancements in AI enable you to integrate and analyze these fragmented data streams efficiently to support a more complete understanding of therapeutics and patient care.
AWS provides a unified environment for multimodal biological foundation models (BioFMs), enabling you to make more confident, timely decision-making in personalized medicine. This AI system combines biological data, model development, scalable compute, and partner tools to support the drug development life cycle. In this post, we’ll explore how multimodal BioFMs work, showcase real-world applications in drug discovery and clinical development, and contextualize how AWS enables organizations to build and deploy multimodal BioFMs.
Multimodal biological foundation models
Biological foundation models (BioFMs) are AI models pre-trained on large biological datasets. BioFMs demonstrate advanced capabilities on specific healthcare and life sciences tasks. The commonly used BioFMs span drug discovery and clinical development domains, particularly in protein structure and molecule design (~20%), omics data analysis including DNA, epigenetic, and RNA (~30%), medical imaging (15%), and clinical documentation (~35%) (Delile et al. 2025).
Unimodal BioFMs are trained exclusively on a single data modality (for example, amino acid sequences) for relevant downstream applications like predicting protein structures; this breakthrough earned the 2024 Nobel Prize in Chemistry. Multimodal BioFMs train across multiple data types (text, audio, image, and video, hereafter “modalities”) and can simultaneously infer across different streams in a single model (for example, text prompts to generate new images or match images to captions).
Notable multimodal BioFM examples include:
- Latent Labs’ Latent-X1 and Latent-X2 not only predict 3D structures of proteins, but also generate novel binders like antibodies, macrocyclic peptides, and miniproteins and predict how they interact with targets.
- Arc Institute’s Evo 2 maps the central dogma of biology to interpret and predict the structure and function of DNA, RNA, and proteins.
- Insilco Medicine’s Nach01 integrates natural language, chemical intelligence, and 3D molecular structure data to accelerate drug discovery.
- Bioptimus’ M-Optimus decodes histology and clinical data for rich biological insights, supporting multiple stages from research to patient care.
- Harvard and AstraZeneca’s MADRIGAL integrates structural, pathway, cell viability, and transcriptomic data to predict drug combination clinical outcome, identify adverse interactions, and optimize polypharmacy management.
- John Snow Lab’s vision language model Medical VLM-24B processes clinical notes, lab reports, and imaging (X‑ray, MRI, CT) for unified, context‑aware diagnostics.
- GEHC’s 3D magnetic resonance imaging (MRI) foundation model, designed to enable developers to build applications for tasks such as image retrieval, classification, image segmentation, and report generation.
The multimodal advantage
The current frontier of models pushes the boundary of multimodal understanding and generation capabilities. General-purpose models like Amazon Nova 2 Omni can process text, images, video, and speech inputs while generating both text and images. This multimodality trend extends to BioFMs, where combining multiple data types like medical images and clinical documentation achieves higher predictive accuracy and broader applicability across diverse clinical outcomes (Siam et al. 2025).
Integrating diverse biological data types yields measurable performance gains:
- Enhanced diagnostic accuracy: Models integrating genomics, imaging, and clinical data yield 4-7% average gains in area under the curve (AUC) over unimodal baselines for diagnoses (e.g., Alzheimer’s, brain cancer) and phenotypes (Sun et al. 2024). Moreover, models integrating lab data, patient exercise metrics, and clinical notes during patient screening achieve 92.74% accuracy with 93.21 AUC in cardiovascular risk prediction (Guo and Wu, 2025).
- Targeted therapeutic strategies: You can use models integrating genomic profiles, medical images, and clinical histories to guide selection of effective interventions for individual patients (Parvin et al. 2025). This proves especially impactful for cancer patients where tumor genomics and radiological imaging can facilitate therapeutic decisions like chemotherapy regimens (Restrepo et al. 2023).
- New disease mechanisms: Single-cell multi-omics models show how cancer cells grow and resist treatments inside blood diseases like leukemia, helping physicians improve survival rates by spotting hidden cancer cells, tracking how mutations drive disease progression, and selecting personalized treatments for patients (Kim and Takahashi, 2025).
- Accurate risk prediction: You can use models integrating lab results, medications, clinical notes, and discharge summaries and other clinical data to predict 30-day hospital readmission risk with 76% accuracy—delivering ~$3.4 million in net savings per hospital annually while improving overall clinical outcomes for high-risk heart failure patients through targeted interventions (Golas et al. 2018).
- Predictive, Preventative, Personalized, Participatory (P4) medicine: Models combining wearable health technologies with patient health data can extract target signals with 96-97% accuracy for diabetes and heart disease diagnosis (Mansour et al. 2021).
BioFMs in action at AWS customers
These performance gains explain why leading biopharma organizations are increasingly adopting multimodal BioFMs. Leading biopharma organizations invest in BioFMs for analyzing biologic (Merck and Novo Nordisk), genomic (AstraZeneca), pathology (Bayer), and clinical (Roche) data. You can realize up to 50% in cost and time savings for drug development and up to 90% in time savings for medical image diagnosis when using these specialized AI models (State of the Art-ificial Intelligence 2025, Jeong et al. 2025). Multimodal BioFMs show promise in multiple stages of the healthcare and life sciences value chain (Figure 1).
Figure 1. Multimodal BioFMs integrate various biological data types (for example, protein, small molecule, omics, imaging, sensors, clinical documentation) to power applications across the drug development lifecycle (research, clinical development, manufacturing, commercial).
For a deeper dive, we’ve selected two use cases: drug discovery and clinical development.
- Designing therapeutic proteins for undruggable disease targets. Multimodal BioFMs integrating computational predictions, structural biology, and biophysical validation enable new approaches to previously inaccessible protein targets (Figure 2). Early applications predicted 3D structures but struggled with multidomain targets featuring discontinuous epitopes. Advanced drug discovery now integrates iterative design-make-test-analyze (DMTA) loops that span structural, computational, and biophysical data. The 3D protein structural data captured through cryo-electron microscopy (Cryo-EM) is evaluated alongside computational metrics like interface predicted template modeling score (iPTM), interface predicted aligned error (iPAE), and root mean square deviation (RMSD) then validated against biophysical measurements such as dose-response curves, biolayer interferometry (BLI), and enzyme-linked immunosorbent assay (ELISA) to accelerate and de-risk drug discovery. For example, Onava’s integrated “AI-human-wet lab” loop represents a step forward in this space by combining generative AI for de novo protein design with rapid experimental validation through an “epitope expansion” strategy, compressing design-to-validation timelines from months to weeks (Calman et al. bioRxiv 2025). You may develop next-generation biologics using multimodal BioFMs like Latent Labs Latent-X2 and Chai Discovery Chai-2 through AWS services including Amazon Bio Discovery, Amazon SageMaker AI for training generative models, Amazon Elastic Compute Cloud (EC2) for model inference, Amazon Simple Storage Service (Amazon S3) for storing structural and experimental data, Amazon Elastic File System (EFS) for shared design libraries, and Amazon Virtual Private Cloud (VPC) for secure infrastructure.
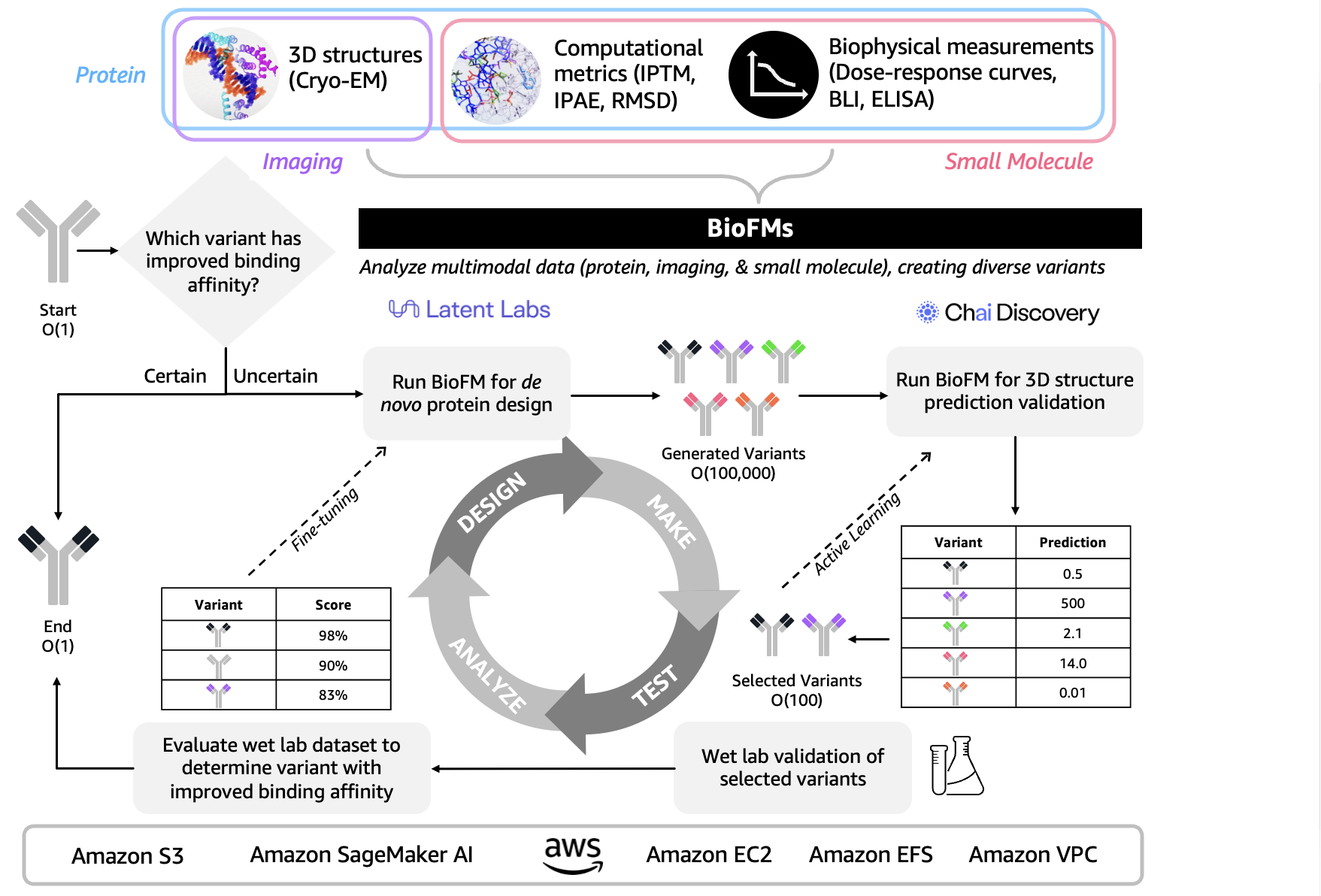
Figure 2. Multimodal BioFMs integrate 3D protein structure, computational metrics, and biophysical measurements through iterative design-validation loops to accelerate therapeutic protein discovery for undruggable multidomain disease targets.
- Predicting immunotherapy resistance in cancer patients during clinical development. Multimodal BioFM developers work towards addressing oncology’s 90% clinical trial failure rate. Today’s multimodal BioFMs simulate tumor microenvironments by integrating sequencing, single-cell data, spatial biology, and patient records to discover resistance mechanisms that reduce patient drop-offs from ineffective treatments and discover new therapeutic targets for previously untreatable patient subgroups (Figure 3). For example, Noetik’s Oncology Counterfactual Therapeutics Oracle (OCTO) simulated 873,000 virtual immune cells across 1,399 patient tumors and revealed why lung cancer patients with KRAS and STK11 gene mutations develop “immune cold” environments blocking immunotherapy effectiveness (Xie et al. Poster presented at SITC 2025). Notably, Noetik achieved 40% faster training time and doubled processing speed through Amazon SageMaker HyperPod’s fault-tolerant infrastructure on AWS with NVIDIA H100 GPUs. You can build your own multimodal BioFMs can take a similar approach using Amazon SageMaker HyperPod for distributed AI training across GPUs, Amazon Elastic Compute Cloud (EC2) for compute capacity, Amazon Simple Storage Service (Amazon S3) for data storage, and Amazon Athena for analyzing petabytes of patient data.
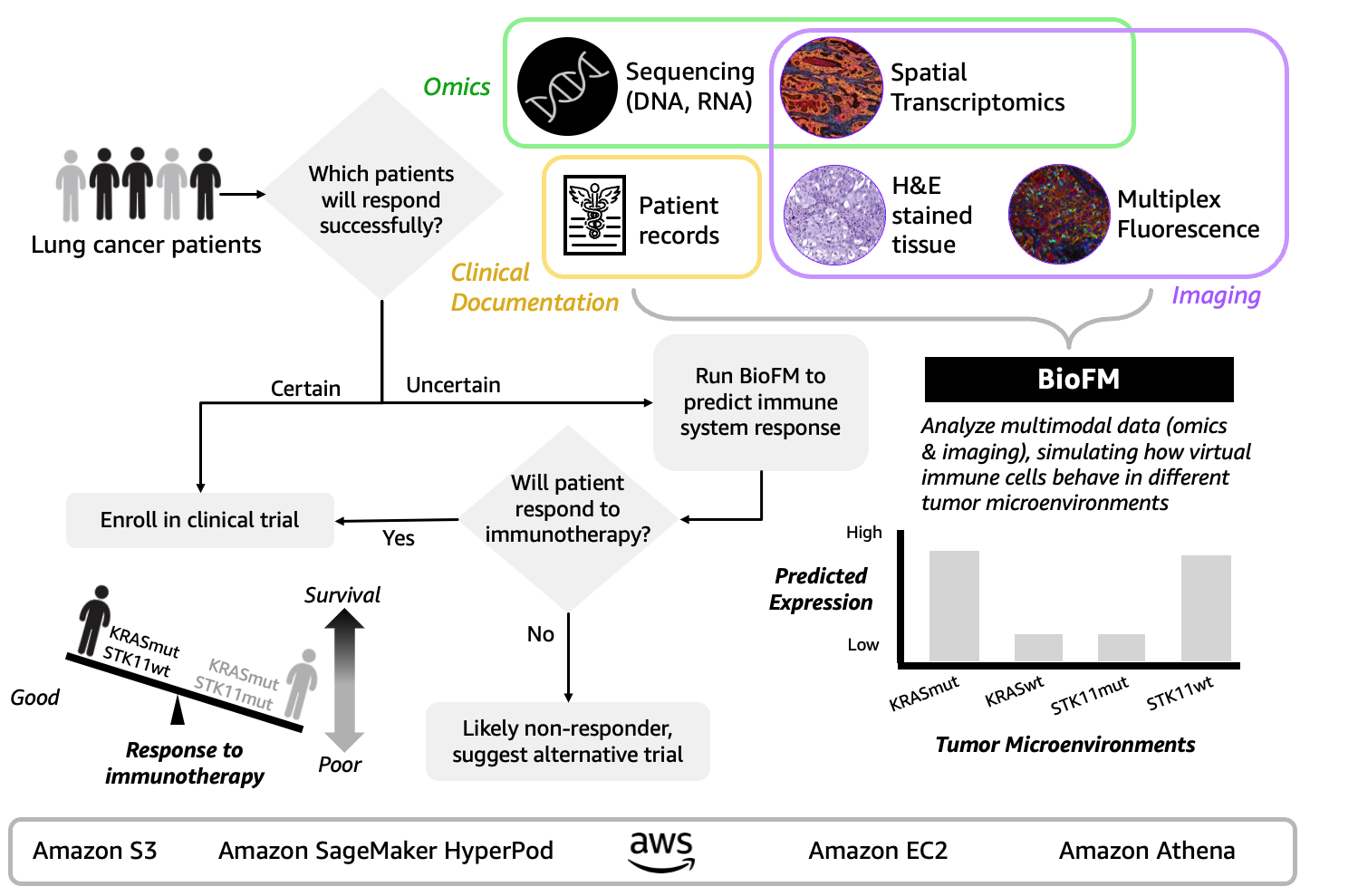
Figure 3. Multimodal BioFM approach combines sequencing, spatial transcriptomics, pathology, and patient records to simulate tumor microenvironments and prioritize patient subpopulations, potentially reducing early-phase trial failures
Solution: AWS environment for multimodal BioFMs
AWS provides a unified environment for building, training, and deploying multimodal BioFMs that help you convert healthcare and life science data into actionable insights. This environment comprises four layers: an AI solution for model development, a unified data foundation for biological data management, scalable infrastructure for compute and storage, and partner integrations that extend capabilities across the drug development lifecycle.
- AI System
- Amazon Bio Discovery provides scientists direct access AI agents selecting the right BioFMs, optimizing inputs, evaluating candidates, sending to lab partners for testing, and automatically returning results for refinement in a lab-in-the-loop cycle that builds institutional knowledge.
- Amazon SageMaker HyperPod delivers distributed training infrastructure for large-scale models. Amazon SageMaker AI compliments this with built-in explainability tools, bias detection, and comprehensive audit trails to support regulatory confidence needed from model development through production deployment.
- Amazon Nova Forge, released at AWS re:Invent 2025, uses the Amazon Nova model family as a starting point to train at optimal points to maximize proprietary data set learning while minimizing training and continued pretraining.
- Amazon Bedrock AgentCore includes the Runtime service to host long-running deep research agents and the Gateway service to securely connect agents to BioFM models and other domain-specific tools.
- Unified Data Foundation
- AWS HealthOmics can orchestrate multi-step AI workflows and handle omics data (DNA, RNA, proteomics) at the petabyte scale, serving as a biological data backbone that powers multimodal BioFM workflows.
- AWS HealthLake and AWS HealthImaging aggregate heterogeneous data into governed lakehouses, automating harmonization across clinical records and medical imaging (radiology, pathology).
- AWS Data Exchange and AWS Lake Formation provide “search, shop, serve” access to federated datasets from Epic, Snowflake, and proprietary sources – revealing disease mechanisms across cancer, rare diseases, and clinical trials without manual integration. AWS Clean Rooms enable federated learning while maintaining data sovereignty.
- Scalable Infrastructure
AWS Partner solutions and implementation support
You can deploy pre-built multimodal BioFMs from partners like NVIDIA directly through AWS. Combine these production-ready NVIDIA NIM microservices with AWS HIPAA-eligible imaging services, multimodal reasoning capabilities, and parallel genomics pipelines to build end-to-end discovery-to-clinic applications. Example partner multimodal BioFMs include:
- MONAI Multimodal: Models combine diverse healthcare data—including CT, MRI, X-ray, ultrasound, EHRs, clinical documentation, DICOM standards, video streams, and whole slide imaging—to enable multimodal analysis for researchers and developers.
- NVIDIA Cosmos: Large Multimodal Models for Science and Medicine. Models like NVIDIA Cosmos Reason-1-7B could be used for surgical robotics training by generating synthetic datasets that combine 3D anatomical models, physics-based sensor data (ultrasound/RGB cameras), and procedural variation.
- La-Proteina: Uses both protein sequence and atom-level 3D structural information to design large, precise proteins, so it can reasonably be described as a multimodal protein model (sequence + structure).
You can consult with implementation partners like Loka, Deloitte, and Accenture on transitioning from proof-of-concept to production deployment for multimodal BioFMs use cases. These partners bring specialized expertise in bioinformatics, cloud architecture, and regulatory compliance to accelerate time-to-value. Visit the AWS Partner Network to explore additional qualified partners with healthcare and life sciences competencies.
Conclusion
Multimodal BioFMs are reimagining what we can discover about disease, treatment, and human health. By integrating omics data, medical imaging, and clinical information, these models reveal hidden insights that were previously difficult to detect through traditional methods. Decision makers can now make more accurate, confident decisions across disease diagnosis, treatment prediction, and therapeutic optimization.
AWS provides a unified environment to overcome the technical barriers of building and deploying multimodal BioFMs at scale. Rather than investing in fragmented, single-use AI solutions for each therapeutic area or clinical application, you can leverage reusable foundation models that adapt across therapeutics and patient care. This system reduces time-to-value while preserving the flexibility to adapt as new data sources and use cases emerge for multimodal BioFMs across therapeutics and patient care.
To learn more about using AWS for BioFM training or inference in a therapeutic or medical context, please contact an AWS Life Sciences representative.
Further reading
About the authors



